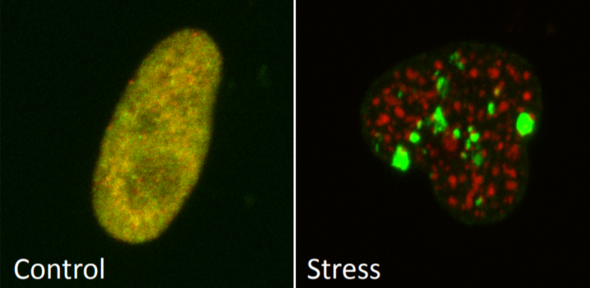
Chromatin control of environmental stress response
Keywords:
Stress, Chromatin, Transcription, Proteostasis, Chaperones
Research Interests:
Cells respond to environmental stress by mounting an adaptive stress response in order to survive stressful conditions. Transcriptional control is a major regulatory layer that determines the strength, the duration and persistence of cellular stress response. Transcription factors, chromatin modifications and non-coding RNA influence the transcriptional response to environmental stress. The molecular mechanisms by which chromatin exerts control over stress response is the main focus of the Unit programme. We aim to address the following questions:
(i) Which cellular pathways sense environmental stress/ toxins and signal to the genome?
(ii) How does chromatin interpret the information about cellular health and toxic exposure determining the transcriptional response to stress?
(iii) How does the transcriptional response adapt cellular phenotypes to survive the stress?
We study these three questions in the context of cellular exposure to environmental stress as well as small-molecule therapeutics in collaboration with pharmaceutical companies. Our approaches include genomics, single-cell transcriptomics, proteomics, chromatin biochemistry as well as genome-wide screening to identify novel components of stress-response pathways. Discovery-driven global approaches in mammalian cells are further validated by in vitro reconstitution experiments and mouse genetic models. We aim to gain novel insights and mechanistic understanding of transcriptional response to stress and toxins.
