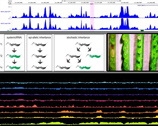
For over a century, research in the Department has explored basic questions in life science from a genetic perspective. While model systems and technologies have changed over this time, central questions on how genotype contributes to phenotype, and how evolution shapes genome function, have been at the core of our work. Today the Department hosts some 27 research groups. centred around genetic approaches to fundamental questions in modern biology. We are an interactive and outlooking department, most of our research groups collaborate with other researchers in the School of Biology, more widely across the University, and with researchers at other institutions.
Developmental Biology
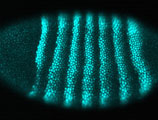

Starting from pioneering research on Drosophila as a model system by John Thoday and Michael Ashburner and colleagues, the Department has a number of Drosophila groups asking fundamental questions in developmental biology [Russell] [St. Johnston] [Karam Teixeira], as part of a wider community in Cambridge using flies as a valuable model system in basic and applied biology. Along with the fly, an expanding repertoire of model organisms is used to explore developmental processes, including plants [Moyroud], mice [Ferguson-Smith] [Imbeault], zebrafish and chick [Steventon], and C. elegans [Ahringer]. As well as tackling developmental questions from the perspective of genes and the genome, the field is becoming increasingly cellular in its focus and analytical methods, with imaging taking centre stage in many research groups [Steventon] [St Johnston] [Clark] [Moyroud]. Key research questions include the roles of cell polarity, signalling, and gene regulation, in cell fate assignments and pattern formation in Drosophila [St Johnston] [Clark] [Karam Teixeira] [Russell], C. elegans [Ahringer], in cultured stem cells or in vivo vertebrate systems [Steventon] [Ferguson-Smith] [Imbeault], and in flower petals [Moyroud].
Cell Biology
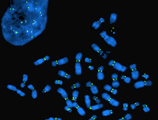
Genetics has catalysed major advances in understanding cell functions. One topic of major interest in the Department is the cell division cycle, including chromosome biology [Farr], asymmetric cell division in yeast [Segal], cell polarity in yeast [Geymonat], and cell polarity and cytoskeleton in Drosophila [St Johnston]. Our work includes regulation and protection of the germline against damage in Drosophila [Karam Teixeira]. In Drosophila neuroscience, interests include circuit approaches to learning and memory and the function and neurological disease roles of the organelle axonal endoplasmic reticulum [O’Kane].
Epigenetic Inheritance
Epigenetic modifications to DNA and chromatin can regulate genome function, host defense mechanisms, and have a profound impact on phenotype. Using a range of approaches and models (computational modeling, environmental responses, imprinting, transposons, RNA-dependent silencing, stem cells, and cancer) and model organisms including plants, Drosophila, C. elegans and mouse, research focuses on the relationships between genotype, epigenotype, post-transcriptional RNA-mediated functions, and normal and abnormal phenotype. In addition, we consider the contribution of dynamic changes in epigenetic state to the properties and programming of stem cells in vivo and in vitro, and the mechanisms regulating the epigenetic programme both within and between generations. Research groups in this area conduct both computational and experimental research and include [Ahringer] [Collepardo] [Ferguson-Smith] [Imbeault] [Sawarkar].
Microbial Genetics
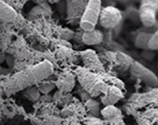

The Department hosts three microbial research groups and two academic visitors. Two groups work on bacterial antibiotic resistance: the role of indole signalling in the formation of biofilms and antibiotic persisters by uropathogenic E. coli , complemented by the work of visitor Claudio [Köser] on regulatory aspects of antibiotic resistance in tuberculosis. Another academic visitor is John [Archer], Founder and Director of HutanBio which applies biology and engineering to develop advanced biofuel sources from natural marine algae. Two groups work on budding yeast, on spindle morphogenesis and asymmetry [Segal] and cell polarity, spindle orientation and cell cycle progression [Geymonat], and together have shown how CDK links cell cycle progression to spindle pole body asymmetry.
Evolution and Population Genetics
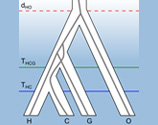 The genomes of living things contain a wealth of information about their evolutionary history, enabling us to infer the causes and timescales of past evolutionary change. Evolutionary research in the Department has a long tradition, and today it combines field and experimental approaches with mathematical and computational techniques, and focuses on a variety of questions, including the co-evolution of pathogens and their hosts [Jiggins], the spread and control of pathogens [Salje], the evolutionary origins of humans and primates [Scally], the genomics of cichlid fish speciation [Durbin], reproductive isolation, and evolutionary inferences from pathogen genomes [Welch]. A common focus of the evolutionary labs is understanding the role of Darwinian natural selection in shaping genomes.
The genomes of living things contain a wealth of information about their evolutionary history, enabling us to infer the causes and timescales of past evolutionary change. Evolutionary research in the Department has a long tradition, and today it combines field and experimental approaches with mathematical and computational techniques, and focuses on a variety of questions, including the co-evolution of pathogens and their hosts [Jiggins], the spread and control of pathogens [Salje], the evolutionary origins of humans and primates [Scally], the genomics of cichlid fish speciation [Durbin], reproductive isolation, and evolutionary inferences from pathogen genomes [Welch]. A common focus of the evolutionary labs is understanding the role of Darwinian natural selection in shaping genomes.
Functional Genomics
 The post-genomic era has stimulated the use of high-throughput techniques for data generation and analyses. The Department is at the forefront of applying genome-wide techniques, including functional and computational genomics, bioinformatics, and genetic databases, to understand and quantify genomic processes in both invertebrate and vertebrate cells. This includes development of algorithms and software for high throughput sequencing data [Durbin] In addition, we are involved in the provision of services to the research community, including InterMine/FlyMine [Micklem], and a state-of-the-art Drosophila facility supporting stock maintenance and transgenesis. Several groups have developed tools for in vivo functional genomics, for example for genome-wide RNAi screening of C. elegans [Ahringer], and genetic resources for the Drosophila community [Russell] [St. Johnston]. Training is increasingly important in the postgenomic world, and the Department has for many years hosted the School of Biological Sciences Bioinformatics Training Facility.
The post-genomic era has stimulated the use of high-throughput techniques for data generation and analyses. The Department is at the forefront of applying genome-wide techniques, including functional and computational genomics, bioinformatics, and genetic databases, to understand and quantify genomic processes in both invertebrate and vertebrate cells. This includes development of algorithms and software for high throughput sequencing data [Durbin] In addition, we are involved in the provision of services to the research community, including InterMine/FlyMine [Micklem], and a state-of-the-art Drosophila facility supporting stock maintenance and transgenesis. Several groups have developed tools for in vivo functional genomics, for example for genome-wide RNAi screening of C. elegans [Ahringer], and genetic resources for the Drosophila community [Russell] [St. Johnston]. Training is increasingly important in the postgenomic world, and the Department has for many years hosted the School of Biological Sciences Bioinformatics Training Facility.
A full alphabetical list of groups in the Department is available HERE